Introduction ¶
BioStudio offers curated, ready-to-run notebooks, multi-omics data, and specialized applications designed specifically for bioinformaticians.
Overview ¶
BioStudio is a powerful tool from BioTuring that enables scientists to analyze biomedical data in a customizable way. With its numerous features and ready-to-use notebooks, users can easily execute and explore their data.
BioStudio offers a variety of tools to help users post their data and analyze the reports. It supports various scientific programming languages such as Python, R, Golang, Julia, Scala, GNU Octave, and Rust, making it ideal for data analysis and education.
Technically, BioStudio is a hosted Jupyter Notebook service that requires no setup, allowing users to quickly build their own notebooks based on their requirements.
Features ¶
Notebook: ¶
-
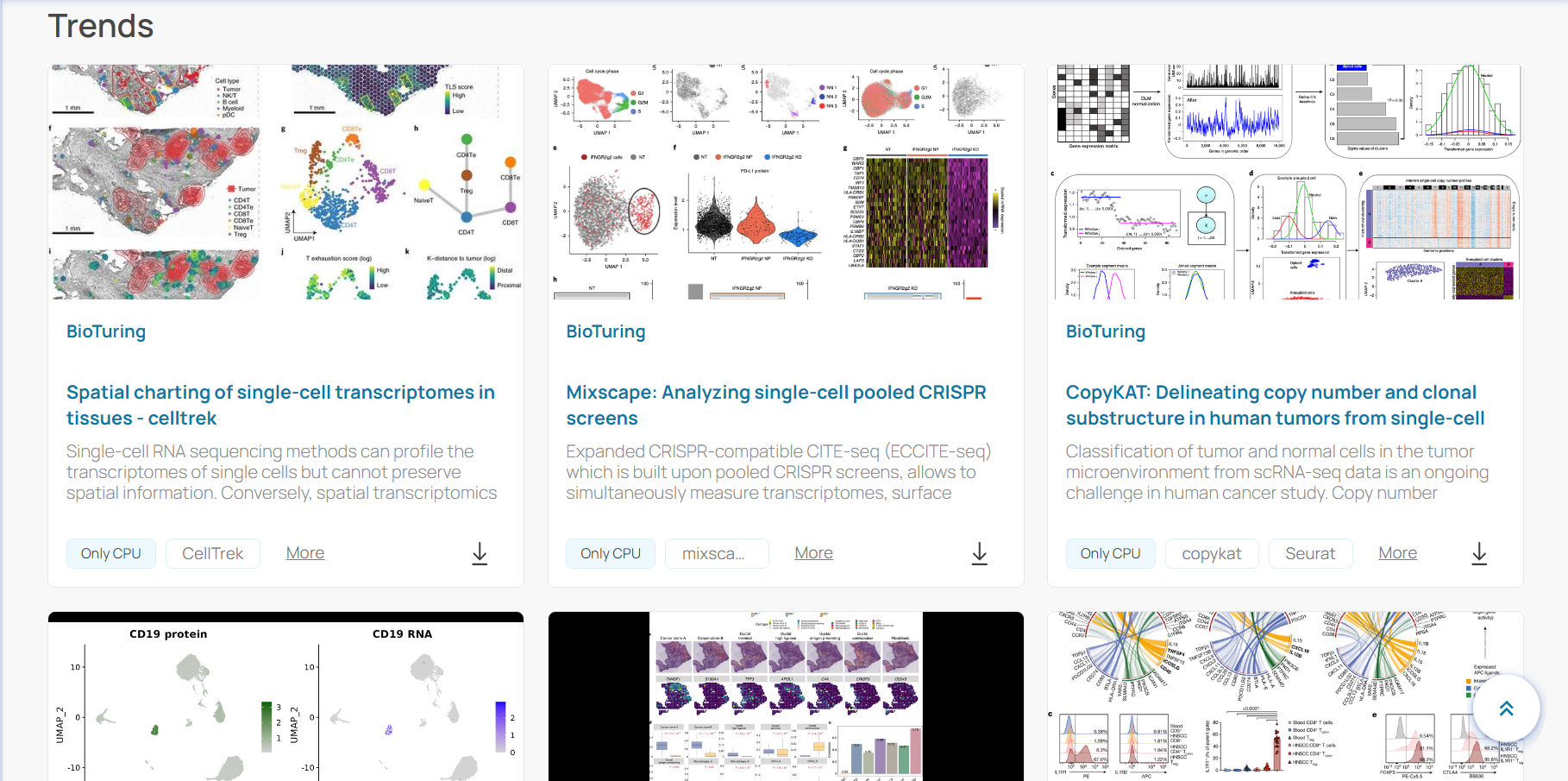
BioStudio provides many ready-to-use notebooks with fully pre-installed environments and executable codes. You just need to download an interesting notebook and run on either example datasets or your own data with a simple click.
-
The notebooks are updated frequently with the latest methods, algorithms, and pipelines, coming with clear instructions and interpretation of example results.
Programming Language: ¶
- BioStudio supports users with a wide range of popular programming languages including R, python, go, Julia, Scala, Octave, Rust, etc.
Package: ¶
-
BioStudio provides pre-prepared library packages, saving users time and effort required for installation.
-
Users can easily create and manage Conda environments using these packages.
-
BioStudio also supports users in creating notebook kernels from their Conda environments.
Study: ¶
-
The BioTuring database currently contains more than 100 million cells from thousands of publications with comprehensive annotation, stringent quality control, and continuous growth.
-
The database contains various multi-omics such as bulk RNA-seq, scRNA-seq, spatial-omics, etc.
-
Once logged in, users can easily search for their desired study and clone the data for further analysis.
Application: ¶
-
BioStudio provides researchers with many interpretative tools for data manipulation in BioStudio workspace such as Shiny apps, Visualization apps, etc. These tools are designed to help researchers easily analyze and interpret their data.
-
Additionally, BioStudio integrates several open-source applications, including Shiny Server, VScode, and File Browsers, into its workspace for enhanced functionality.
Cooperation: ¶
- With BioStudio, users can seamlessly access data from other platforms within the BioTuring ecosystem, such as BBrowserX, Talk2Data, BioTuring Lens, and BioVinci. This feature allows for efficient data sharing and collaboration across multiple research purposes.
Resource efficiency: ¶
- BioStudio optimizes the use of computational resources, including RAM and CPU, to execute multiple tasks efficiently. This feature enables users to perform various analyses without requiring high-performance PCs or excessive computational power. |
Example notebooks ¶
Architecture ¶
The BioStudio architecture diagram provides a high-level view of the components and relationships of our system. It shows how our different services and systems interact with each other and with all services.
Installation ¶
BioStudio provides 2 ways to install:
On private server. ¶
-
It is installed and runs on computers on the premises of the person or organization using the software, rather than at a remote facility such as a server farm or cloud.
-
For more details, checkout this link: Installation Steps
On BioTuring cloud. ¶
-
It is the on-demand availability of computer system resources, especially data storage (cloud storage) and computing power, without direct active management by the user.
-
For more details, contact us: support@bioturing.com
